High-resolution immune cell atlases reveal how inherited DNA and lifelong environmental exposures leave distinct epigenetic signatures that together program immune function and disease risk.
study: Genetics and environment uniquely shape the epigenome of human immune cells. Image credit: Anusorn Nakdee / Shutterstock
In a recent study published in natural geneticsresearchers have created a comprehensive atlas to elucidate the combined effects of genetic inheritance and environmental exposures on the human immune system. The study analyzed blood samples from 110 donors who had been exposed to pathogens such as HIV or COVID-19, or chemicals such as pesticides, and found that environmental and genetic factors targeted different parts of the genome.
Specifically, genetic factors primarily affect the gene bodies of memory immune cells, whereas environmental factors tend to fine-tune the regulatory “switches”, enhancers, and promoters of naive immune cells. These findings provide a high-resolution map of how infection history and underlying DNA combine to shape immune cell state and function relevant to human health, and epigenetic changes frequently coincide with changes in chromatin accessibility in regulatory regions.
Background: Nature, nurture, and immune plasticity
Conventional wisdom held that environmental exposures determined an individual’s long-term health outcomes. With the discovery of genetics and DNA, this wisdom moved into a debate over the relative importance of nature, genetics, nurture, and environment in human development.
Scientists agree that one of the most important models of the interaction between genetics and environment is the immune system. The immune system must be rigid enough to distinguish between self and non-self, yet flexible enough to learn from new threats.
Previous research has long established that DNA sequences are inherited and cannot be changed. However, more recent studies hypothesize that how these genes are used is controlled by the epigenome. The epigenome is a mechanistic interaction between genetics and environment that results in changes in gene expression without altering the underlying DNA sequence.
Epigenetic regulation and DNA methylation
One of the main tools used by the epigenome is DNA methylation. DNA methylation is the addition of chemical tags, methyl groups, to DNA that act like a dimmer switch, increasing or decreasing gene expression without changing the underlying code. Previous studies have investigated the effects of environmental exposures on epigenetic outcomes, but have primarily focused on bulk tissues, obscuring the details of how specific immune cell types and conditions respond.
Research design and sample population
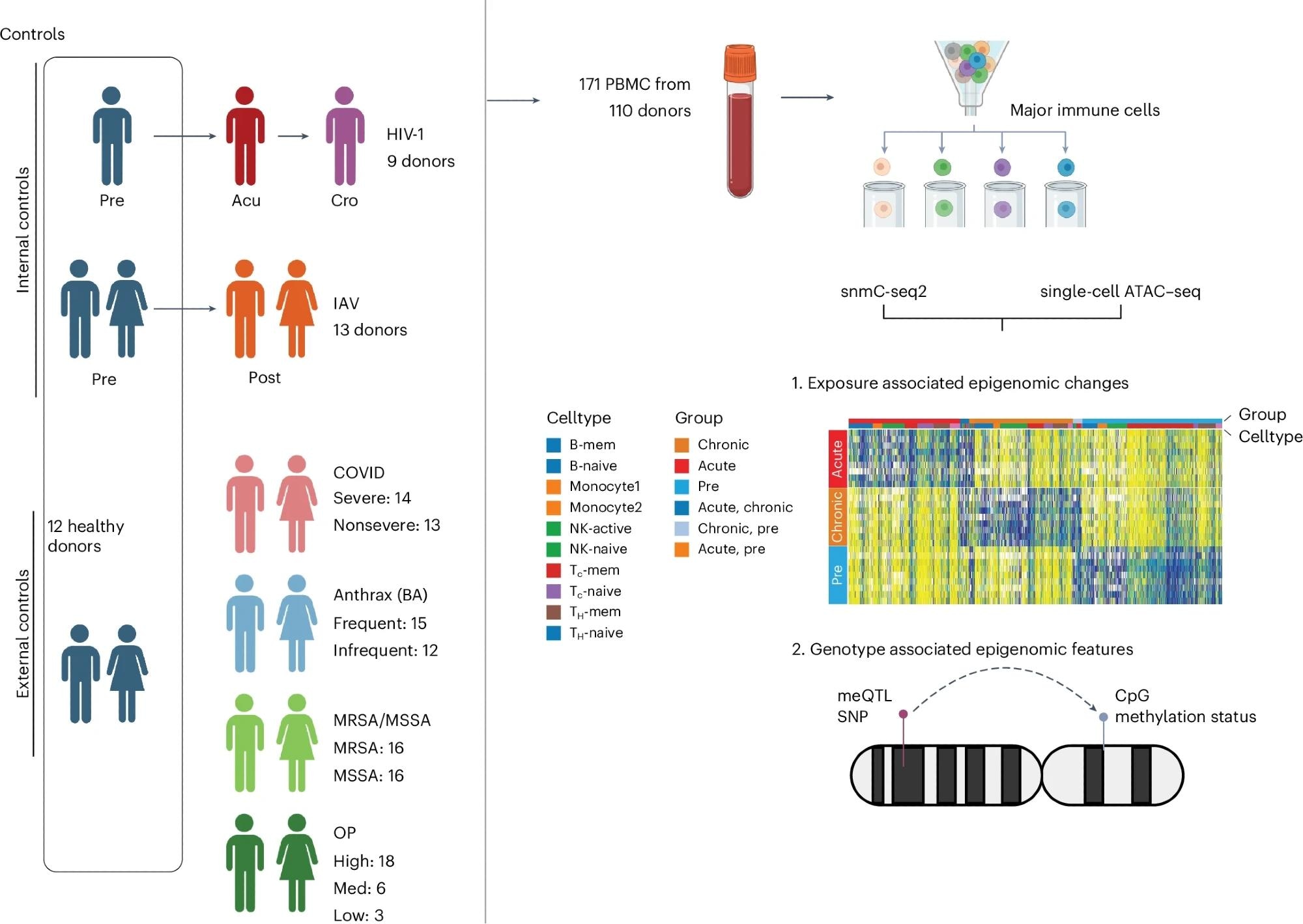
In this study, 171 peripheral blood mononuclear cells (PBMC) Sample from 110 people.
The sample cohort included both healthy donors and individuals with specific defined exposures.
For HIV-1 and IAV, we have internal control samples taken from the same set of donors before infection and samples taken after exposure. We also collected PBMCs from 12 healthy donors as external controls. For exposures without internal controls (COVID-19, anthrax vaccine, MRSA/MSSA, OP), all healthy samples were used as controls. We performed snATAC-seq and snmC-seq2 on PBMCs to identify eDMRs associated with exposure and genotype. We also used this dataset to identify gDMRs. BA, anthrax; The diagram is created as follows BioRender.com.
Types of environmental and biological exposure
-
Viral infections such as HIV-1 and influenza A (IAV), and SARS-CoV-2 (COVID-19).
-
Bacterial infections such as methicillin-resistant bacteria Staphylococcus aureus (MRSA) and MSSA.
-
Exposure to chemicals or vaccines, especially organophosphate pesticides (OP) and anthrax vaccine.
Experimental methods and cell type isolation
Experimental assays included the preparation of an snmC-seq2 library, which enabled sequencing of DNA methylation in single cell nuclei. Participants’ blood samples were categorized into seven major immune cell types: T cellsB cells, natural killer (N.K.) cells, and monocytes. In downstream analyses, we further disaggregated these populations into naïve and memory lymphocyte states as well as additional immune subtypes.
Longitudinal and comparative analysis
Longitudinal internal controls were available for exposure to HIV-1 and influenza A viruses, and samples were collected from donors before and after infection. Other exposure analyzes relied on external controls and statistical adjustments, and thus were more limited in their ability to capture within-individual epigenetic changes.
By comparing these groups to controls, statistical analyzes in this study sought to elucidate regions that were differentially methylated in relation to exposure (eDMR) and differentially methylated regions associated with genotype (gDMR).
Epigenetic signatures of exposure and inheritance
-
eDMRexposure-related differentially methylated regions, changes associated with environmental exposure.
-
gDMRdifferentially methylated regions associated with genotype, changes associated with genetic genetic variation.
Key findings: Genome-wide epigenetic mapping
snmC-seq2 analysis in this study identified 756,575 environmental methylation markers (eDMR) and over 275,000 genetic markers (gDMR) across the immune cell types investigated. The most remarkable finding of this analysis was that nature and upbringing appear to function in different “regions” of the genome.
Genomic targets with different genetic and environmental influences
different genomic targets, eDMR Enhancer- and promoter-related regulatory regions, or genomic switches that influence when genes are turned on and off, were consistently enriched. in contrast, gDMR They were mainly located within the gene body. This suggests that while genetic variation shapes the core protein-coding structure, environmental exposures preferentially alter the regulatory networks that control gene activity, often altering regions that exhibit coordinated changes in chromatin accessibility.
Cell state specificity of epigenetic effects
Specificity of cell state, environmental exposures were observed to have a more pronounced effect on naive lymphocytes, whereas genetic factors had a stronger effect on memory lymphocytes, reflecting the cumulative effect of genetic variation on the state of long-lived immune cells.
Immune remodeling in HIV and COVID-19
Impact of COVID-19 and HIV, previous HIV-1 infection was associated with significant changes in immune cell composition, particularly remodeling N.K. cells and memory T cells.
Regarding the novel coronavirus disease (COVID-19), analysis identified the presence of unique monocyte clusters in critically ill patients. The statistical separation of this cluster was substantial (P = 2.05 × 10^-237), suggesting that these cells are strongly associated with the inflammatory immune program characteristic of severe COVID-19 without establishing a direct causative role in disease pathology.
Epigenetic variations associated with genetic ancestry and disease
Ancestry and disease risk, genetic ancestry was found to influence immune-epigenetic responses. Either MRSA Due to COVID-19 and exposure to COVID-19, individuals with African genetic ancestry showed greater methylation changes than individuals of European descent, even after adjusting for covariates.
Additionally, genotype-associated methylation signals overlapped with known disease risk loci. Specifically, genetic variants associated with eczema and cholelithiasis showed significant colocalization with methylation sites on T cells, linking immune-epigenetic regulation to broader disease susceptibility.
Conclusions and implications for personalized medicine
This study provides evidence that nature and nurture are distinct architects of immune cell biology. Although their effects are focused on immune function, genetic variation and environmental exposures appear to act through distinct mechanistic and genomic pathways.
Atlas has the potential to advance personalized medicine. Future studies will be needed to verify the biological and clinical relevance of specific substances. eDMR and gDMRtheir functional impact needs to be clarified and current limitations such as incomplete exposure histories, modest sample sizes in some exposure groups, and limited longitudinal sampling need to be addressed before they can be reliably introduced as high-precision biomarkers.